About Me

Who I am: My hobbies outside of science include spelunking and cuddling with my cat Toki Roki. I became fascinated by research in high school after joining the Zoo School program, which allowed me to complete a 6 month behavioral study on two cougars at the Columbus Zoo and Aquarium.
Education: B.S. in Biology, Specialization in Ecology and Evolution from University of Chicago 2022
Research Interests: I am interested in studying how many environmental yeast species are responding to changes in the environment. I am working on improving our ability to find pathogenic yeast species in the environment in addition to studying what genes appear to be undergoing changes within populations of wild yeast.
Publications: Coming soon!
Research
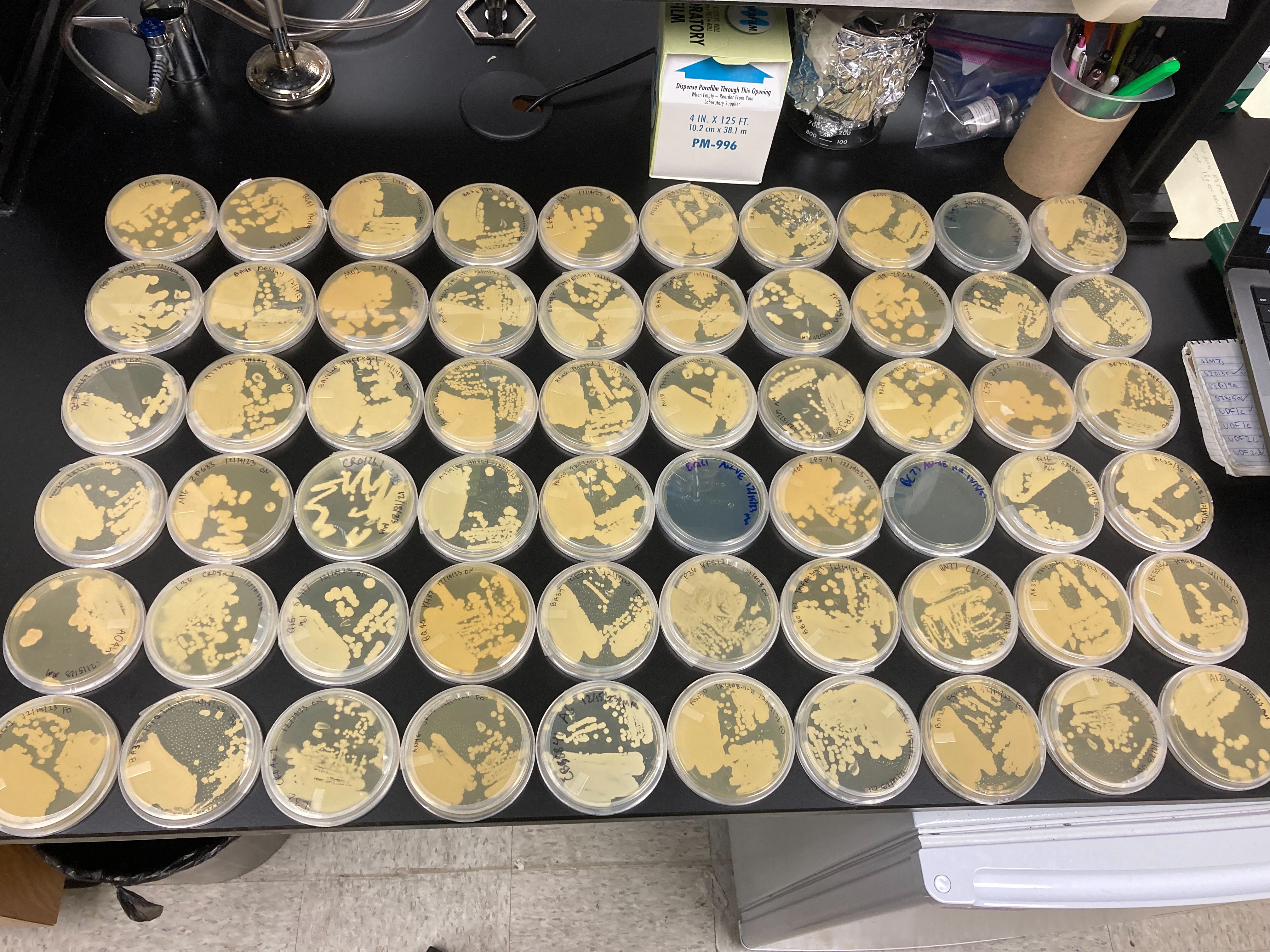 60 plates with yeast streaked.
60 plates with yeast streaked.
Phylogeography and Population Genetics of Saccharomyces paradoxus
I work in the Bensasson lab at UGA, and one of the species I study in my research is a woodland yeast known as S. paradoxus. Our lab has made some surprising discoveries about the range of this species in North America. We added 44 new North American and 46 new European genomes, and our lab was able to detect changes occuring in the DNA of multiple S. paradoxus populations.
Improving isolation of pathogenic yeast from the environment
Another aim of my PhD is to improve the isolation of pathogenic yeast from environmental sources. With funding from the NIH T32 Genetics Training grant I experimented with different types of enrichment media, incubation temperatures, and plate media in an attempt to target pathogens from environmental samples.
Research Relevance
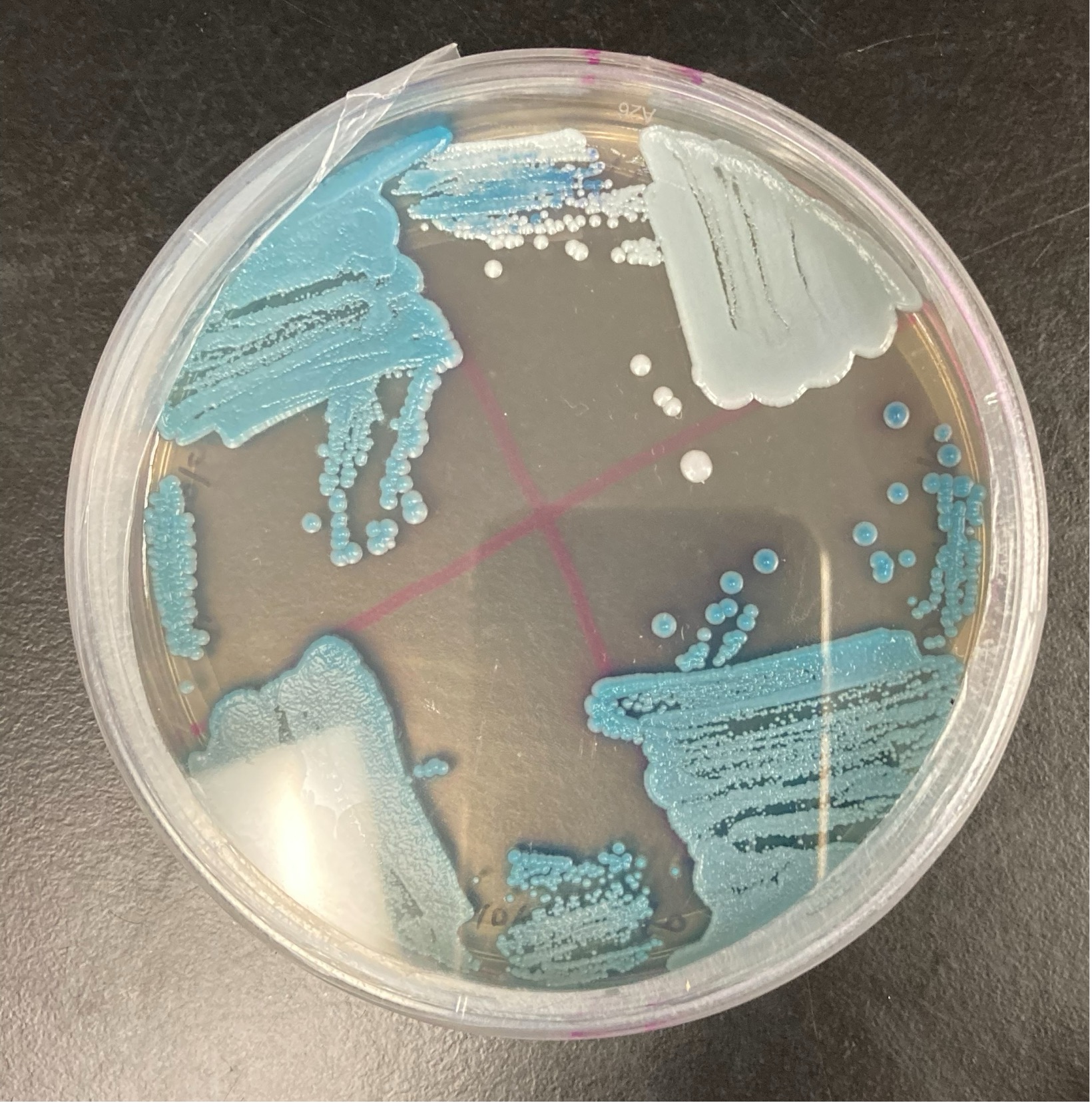 Yeast species in the Candida genus may grow blue on CHROMagar plates.
Yeast species in the Candida genus may grow blue on CHROMagar plates.
Hidden pathogens: Yeast Among Us
While potentially harmful yeast can be found in natural environments around us, there is still a lot we don’t know about them, including their exact locations. The results of my research will give us better tools to find the wild counterparts of yeast strains usually associated with human illness.
A top concern for the World Health Organization are two yeast species in the critical priority pathogens list: Candida albicans and Candida auris. C. albicans is the most common human fungal pathogen, while C. auris was first identified as a pathogen in 2009 but quickly gained international attention. C. auris arose as a pathogen simultaneously in three different continents, shifting from a harmless environmental yeast to a deadly vector of disease as it adapted to higher temperatures, allowing it to survive in the human body. Possibly due to antifungal exposure in agricultural fields, C. auris is also resistant to nearly every known antifungal treatment, contributing to a high mortality rate in patients who contract it. The case of C. auris highlights the urgency of investigating environmental yeast strains, understanding how they are adapting in the face of climate change and how it may impact human health.
But the methods used to identify yeast have a major flaw---When there are multiple yeast species present in a sample, depending on what types of media (food for the yeast) and temperatures researchers use, some yeast may not survive and will never be detected. Current methods mainly target yeast that can be used for bread making or brewing, and the few methods that target pathogens usually target the DNA, identifying where the yeast are but not keeping a living sample that can be used for experiments or studied further. Only 1% of isolated live environmental yeast are pathogens, although DNA methods suggest there are many more.
Despite their prevalence in nature and impact on human health, environmental yeast are understudied. It wasn’t until 2018 that researchers discovered that the yeast that causes oral thrush, C. albicans, had thriving wild populations on oak trees and in soil rather than only living on and inside animals. A factor of the mystery behind the wild populations of pathogenic yeast in the wild is the inability to find these yeast species.
Yeast are competing for space with thousands of other microbes. When we take soil or bark samples looking for yeast, we’re essentially going in blind, and we are only able to find yeast by putting them in laboratory conditions that give yeast the advantage to wipe out their competition and reproduce enough that they form colonies we can see.
The Bensasson lab is developing new enrichment and media methods that target pathogens in the environment. With these new methods, we will hopefully be able to get a better sense of what types of pathogens are in our environments. In the future, when we have more live environmental strains, we will also be able to investigate whether environmental yeast strains and clinical yeast strains (those taken from people) are interacting with one another. This could be very important from a healthcare perspective, as clinical strains could pass on genes to wild strains that enhance their ability to infect humans or vice versa.
In addition to pathogens, I am focusing my research on an environmental yeast species that is not associated with humans, Saccharomyces paradoxus. This species is quickly evolving, and because it is closely related to a well studied species, Saccharomyces cerevisiae, there is already a lot known about the functions of many genes that may be changing to increase the ability of S. paradoxus to survive.
Contact
Email: mmm07490@uga.edu
Address: 2502 Plant Sciences Bldg Athens, GA 30602-0001




